Command Palette
Search for a command to run...
AI-driven De Novo Design of Diverse small-molecule Binding Proteins: A South Korean Team Discovered a Protein That Can Selectively Recognize Stress hormones.

In the fields of life sciences and synthetic biology, designing small-molecule binding proteins with both high affinity and high specificity has always been a key challenge for realizing biosensing and molecular switches. In the past, this approach mainly relied on the screening and modification of natural proteins, or on physical modeling designs based on existing protein backbones, which has always limited versatility and scalability.
In view of this,A research team from the Department of Biological Sciences at the Korea Advanced Institute of Science and Technology (KAIST) has used deep learning-driven protein structure generation and sequence design methods to design diverse small-molecule binding proteins de novo, using the NTF2-like fold as a core "universal backbone."Furthermore, they transformed this into a sensor similar to chemically induced dimerization (CID). Researchers successfully designed a protein capable of selectively recognizing the stress hormone cortisol and developed an artificial intelligence biosensor based on it. This case transcends protein design itself, moving towards practically measurable sensor technology and solving the long-standing challenge of small molecule recognition in the field of protein design.
The related research findings, titled "Small-molecule binding and sensing with a designed protein family," have been published in Nature Communications.
Research highlights:
* Utilizing artificial intelligence to design proteins from scratch (de novo) and apply them to functional biosensors.
Traditional methods primarily involve finding natural proteins or modifying certain functions, while this study uses artificial intelligence-based design to "customize" proteins with the desired functions.
* The research findings can be widely applied in fields such as disease diagnosis, new drug development, and environmental monitoring.

Paper address:
https://www.nature.com/articles/s41467-026-70953-8
More AI frontier papers:
https://hyper.ai/papers
Dataset: Building the NTF2 backbone
To achieve the design goals,The researchers first generated a set of NTF2 structures (set 1: 1,615 backbones) using a family-level "hallucination" method, and then used ProteinMPNN to redesign the sequences of these backbones.The AlphaFold algorithm was used to screen proteins that could fold into the designed structures (set 2: 3,230 backbones). Furthermore, Rosetta parameterization was used to generate the backbone structure, and ProteinMPNN was used for sequence design, followed by AlphaFold for structure validation (set 3: 6,838 backbones), as shown in the figure below:
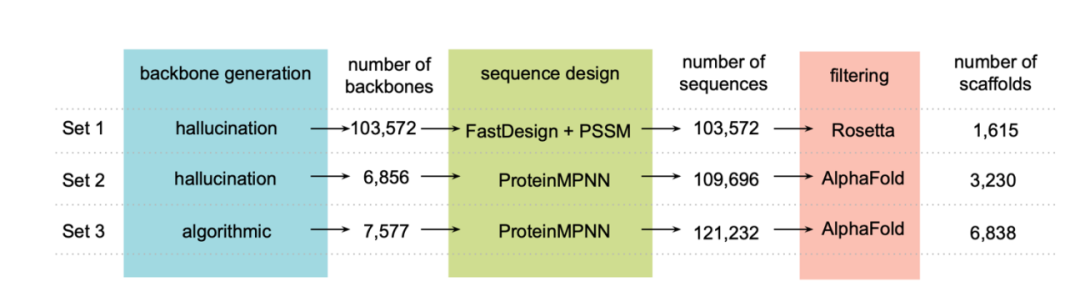
Ultimately, after screening, the researchers also obtained the encoding oligonucleotides for experimental characterization, including: 630 HCY-binding proteins, 1,661 ROC-binding proteins, 16,276 WRF-binding proteins, 9,024 APX-binding proteins, 19,390 IRI-binding proteins, and 7,573 OHP-binding proteins.
Designing the NTF2 protein family with diverse pocket geometries
The NTF2 fold consists of three α-helices and a curved 6-strand β-fold sheet. These structures together form the large internal connecting pocket characteristic of this fold family, as shown in the figure below:
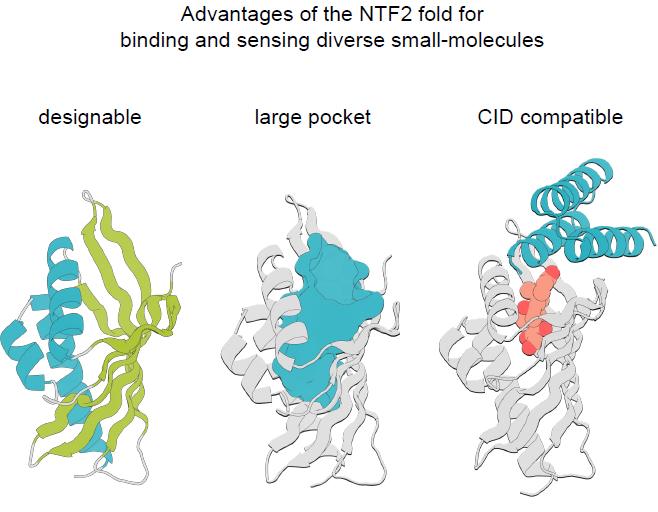
The diversity of this folding in nature mainly stems from the long and irregular rings and the unique quaternary structure, both of which influence the geometry and function of the combined pocket.The goal of this study is to design a family of NTF2 proteins with diverse pocket geometries to accommodate a wide range of small molecules, while minimizing loop regions to maintain their modularity and designability.The overall design process is shown in the following diagram:
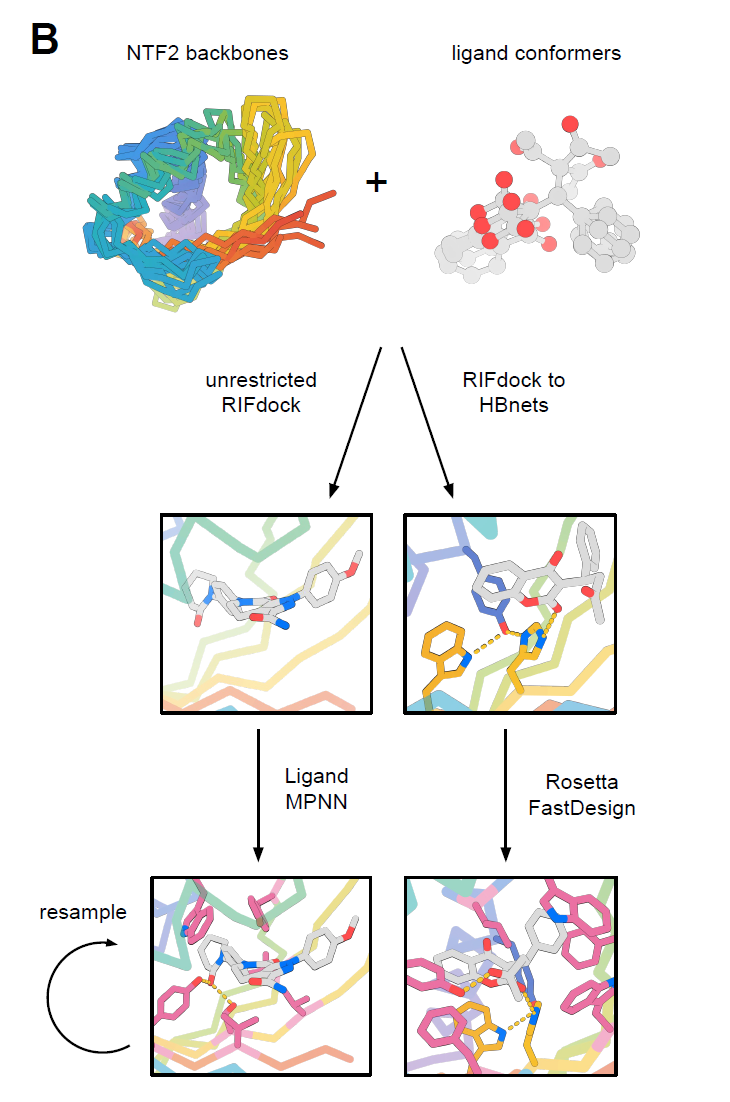
After obtaining more than 10,000 NTF2 designed proteins with diverse pocket geometries, researchers used RIFdock to place six small molecules, which differ in chemical properties and structure, into the central pockets of these backbones. These small molecules included the stress hormone cortisol (HCY), the anticoagulant warfarin (WRF), the muscle relaxant rocuronium bromide (ROC), the anticoagulant apixaban (APX), the antitumor active molecule SN-38 (IRI) derived from the anticancer drug irinotecan, and the hormone 17-α-hydroxyprogesterone (OHP).
Constructing polar interfaces is a significant challenge in protein design, especially for small-molecule binding proteins. This requires introducing polar residues into the internal pocket to interact with the polar functional groups of the ligands, while simultaneously maintaining the overall stability of the protein. To address this, researchers employed two strategies:
Method 1 (RIFdock to HBNets)
Researchers hooked HCY, WRF, ROC, APX, and IRI into the 1 backbone and required at least one protein-small molecule interaction mediated by HBNet residues. They then optimized the design using a Rosetta design guided by natural sequences, which used a position-specific scoring matrix derived from NTF2 family proteins to bias the sequence design.
Method 2 (unrestricted RIFdock)
OHP, APX, and IRI were placed into the backbones of sets 2 and 3 using unconstrained RIFdock, and sequence design was performed using LigandMPNN. LigandMPNN is a variant of ProteinMPNN specifically trained on protein-small molecule complex data, enabling explicit consideration of ligand presence during the design process.
When screening design results, researchers used Rosetta to calculate the number of hydrogen bonds between protein and ligand, binding energy (ddG), and contact molecular surface area (CMS); for method 2, they also combined single-sequence AlphaFold prediction results to screen designs that could simultaneously reproduce the target fold structure and binding site (see figure below).
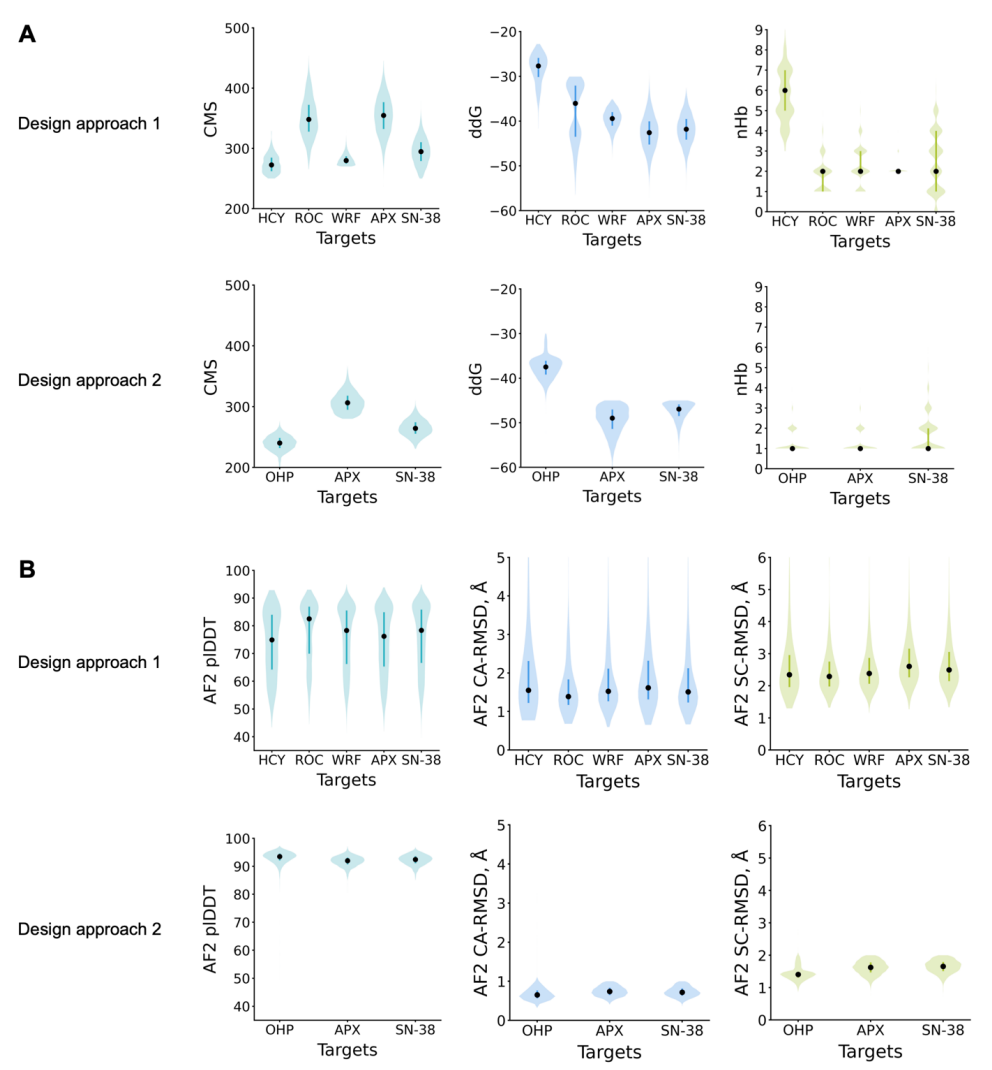
Results Showcase: NTF2-based small molecule binding proteins can be applied to biosensors
Researchers designed a series of experiments to verify the effectiveness of the design strategy proposed in this study:
Design and structural characterization of binding proteins
To verify the accuracy of the designed small-molecule binding proteins, the crystal structures of two protein-ligand complexes were resolved: the cortisol-binding protein hcy129 and the apixaban-binding protein apx1049. Specifically, hcy129 underwent surface redesign using ProteinMPNN to improve crystallinity, successfully obtaining its 1.5 Å high-resolution structure with the cortisol complex. Structural alignment showed that the overall folding was highly consistent with the design model, with a Cα RMSD of 1.1 Å (Figure A below). The key hydrogen bond residues and ligand conformations also matched precisely (Figure B below), indicating that the pre-constructed hydrogen bond network (HBNet) effectively achieved the precise design of polar interactions.
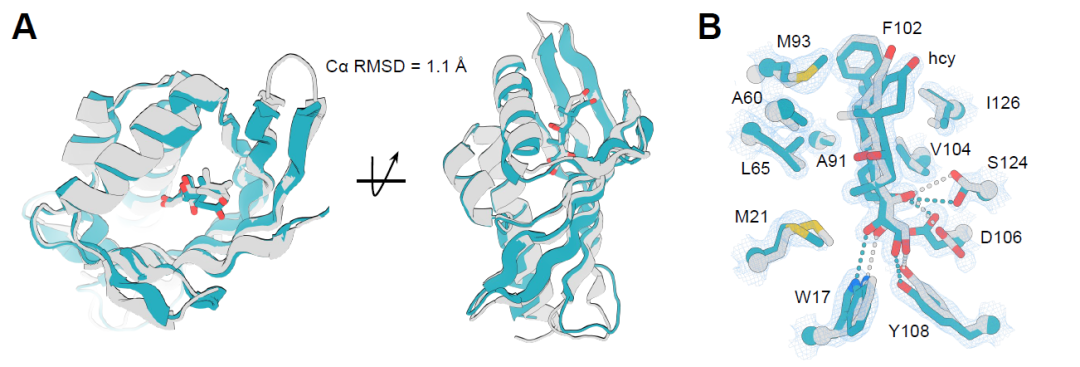
On the other hand, the crystal structure of the apx1049-apixaban complex has a resolution of 2.1 Å, showing higher consistency with the design model, and a Cα RMSD of only 0.6 Å over a range of 113 residues (Figure C below). Its protein-ligand interactions almost perfectly replicate the design, including key hydrogen bonds and π–π stacking interactions between aromatic residues (Figure D below), thereby stabilizing the ligand conformation and forming a highly shape-complementary binding pocket. These results demonstrate that this design strategy achieves high-precision protein-ligand interface construction at the atomic scale.
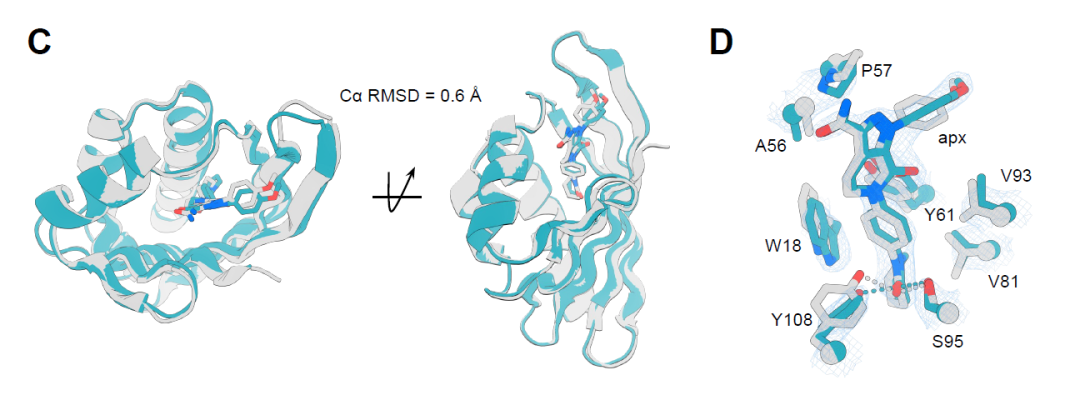
Design of specificity assessment for binding proteins
To evaluate the specificity of the designed proteins, the study systematically tested six binding proteins with six ligands, using albumin, which has non-specific binding ability, as a control. The results showed that high-affinity proteins such as hcy129.1, iri807.1, and apx1049 exhibited good specificity when binding to their respective targets, while albumin showed almost no binding to most ligands, validating the effectiveness of the design strategy.
Furthermore, in the warfarin (WRF) system, the binding affinity of albumin (KD approximately 5.0 μM) to it is similar to that of the designed protein wrf1071 (KD approximately 1.1 μM), indicating that nonspecific binding to highly hydrophobic ligands remains challenging.
Overall, this method has achieved a certain degree of high specificity in recognition, but there is still room for further optimization in distinguishing structurally similar molecules and improving selectivity for hydrophobic ligands.
Biosensor Construction (Design and Characterization of Cortisol-Induced Heterodimers)
Cortisol is typically present in physiological samples at low nanomolar concentrations, but plasma cortisol levels above 38 nM can be used as a diagnostic criterion for diseases such as Cushing's syndrome. To improve the binding affinity of hcy129 for cortisol for biosensing, researchers constructed a combinatorial mutant library based on favorable mutations screened in its single-point saturation mutation (SSM) experiments. Screening was performed using yeast display, and a significant increase in binding affinity was observed, as shown in the figure below.
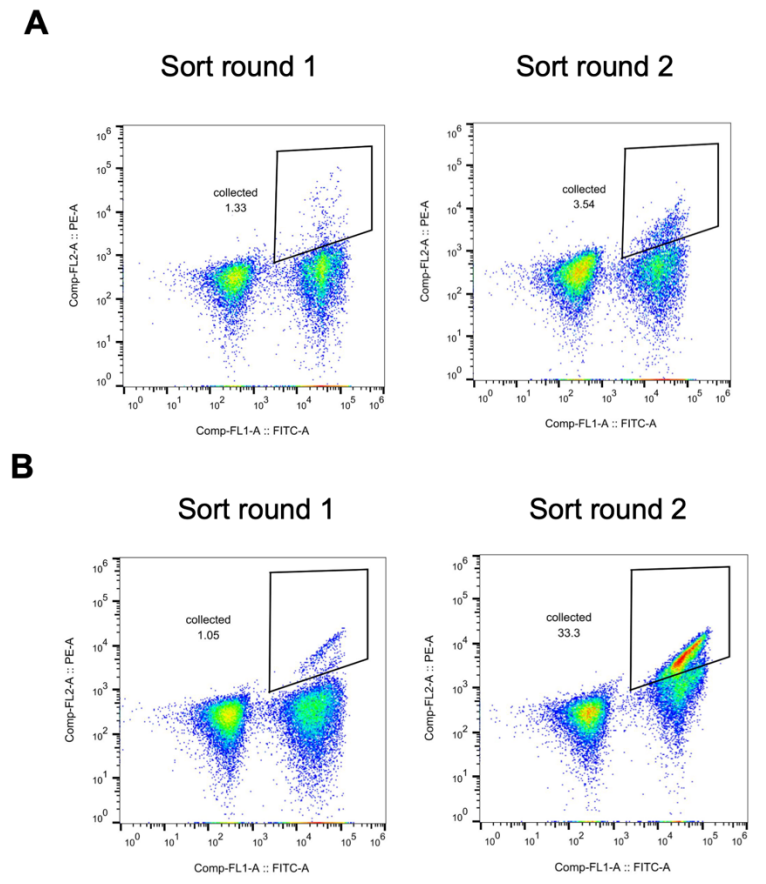
Subsequently, the researchers screened the optimal variant from the library, expressed it in E. coli, and characterized it using isothermal titration calorimetry (ITC). The KD of this variant hcy129.1 was 68 nM, which was 31-fold higher than the original design (Figure C below); structural analysis showed that the enhanced affinity mainly stemmed from a stronger hydrophobic interaction with cortisol (Figure D below).
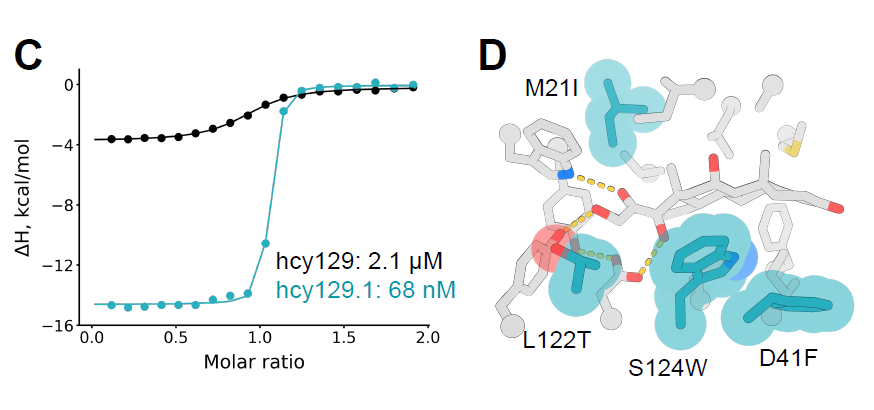
Building upon this foundation, the study further designed a cortisol-dependent heterodimer system. By modifying the structure of hcy129.1 and introducing a small protein backbone, computational design and screening were performed using methods such as RIFdock, Rosetta, and ProteinMPNN. Ultimately, a small protein, miniH11, was obtained that can form a ternary complex with hcy129.1 and cortisol.
Experiments showed that the system forms a stable complex only in the presence of cortisol. Furthermore, the system was fused with a NanoBiT luciferase system to achieve cortisol sensing, detecting an EC50 of approximately 72 nM (Figure H below), consistent with the binding affinity, thus validating the design's effectiveness. Simultaneously, the affinity of the system significantly decreased in the absence of cortisol, indicating that the dimerization process exhibits good ligand dependence.
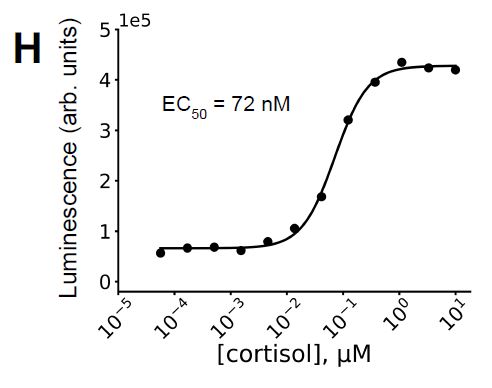
In the equimolar (200 nM) hcy129.1_CID-SmBiT and miniH11-LgBiT system, the cortisol-dependent luminescence response curves...
Overall,This work demonstrates that NTF2-based small molecule binding proteins can be further engineered into functional biosensors.
Conclusion
Overall, this study provides a novel pathway for the de novo design of small molecule binding proteins: by using artificial intelligence models to precisely characterize protein-ligand interactions at the atomic level, it achieves a shift from "discovering or modifying natural proteins" to "customizing functional proteins on demand," and completes effective validation at the experimental level.
This not only marks a leap forward in protein design capabilities but also significantly expands its application boundaries—from precise detection of biomarkers in early disease diagnosis to targeted molecular recognition in new drug development, and real-time perception of pollutants in environmental monitoring. As this technology matures, highly specific and programmable customized biosensors are expected to become a crucial bridge connecting life sciences with real-world applications.
References:
https://www.nature.com/articles/s41467-026-70953-8
https://phys.org/news/2026-04-ai-proteins-built-specific-compounds.html